Abstract
Group 2 innate lymphoid cells (ILC2 cells) are important for type 2 immune responses and are activated by the epithelial cytokines interleukin 33 (IL-33), IL-25 and thymic stromal lymphopoietin (TSLP). Here we demonstrated that IL-1β was a critical activator of ILC2 cells, inducing proliferation and cytokine production and regulating the expression of epithelial cytokine receptors. IL-1β also governed ILC2 plasticity by inducing low expression of the transcription factor T-bet and the cytokine receptor chain IL-12Rβ2, which enabled the conversion of these cells into an ILC1 phenotype in response to IL-12. This transition was marked by an atypical chromatin landscape characterized by the simultaneous transcriptional accessibility of the locus encoding interferon-γ (IFN-γ) and the loci encoding IL-5 and IL-13. Finally, IL-1β potentiated ILC2 activation and plasticity in vivo, and IL-12 acted as the switch that determined an ILC2-versus-ILC1 response. Thus, we have identified a previously unknown role for IL-1β in facilitating ILC2 maturation and plasticity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
Change history
29 April 2016
In the version of this article initially published online, the interferon (IFN-α) in the fourth sentence of the abstract is incorrect. That section should read "the locus encoding interferon-γ (IFN-γ)...." Also, sentence two in paragraph three of the introduction includes a typographical error ("zand"); that should read "IL-18 and IL-1...." The errors have been corrected for the print, PDF and HTML versions of this article.
References
Sonnenberg, G.F. & Artis, D. Innate lymphoid cells in the initiation, regulation and resolution of inflammation. Nat. Med. 21, 698–708 (2015).
Neill, D.R. et al. Nuocytes represent a new innate effector leukocyte that mediates type-2 immunity. Nature 464, 1367–1370 (2010).
Moro, K. et al. Innate production of T(H)2 cytokines by adipose tissue-associated c-Kit+Sca-1+ lymphoid cells. Nature 463, 540–544 (2010).
Price, A.E. et al. Systemically dispersed innate IL-13-expressing cells in type 2 immunity. Proc. Natl. Acad. Sci. USA 107, 11489–11494 (2010).
Tait Wojno, E.D. & Artis, D. Innate lymphoid cells: balancing immunity, inflammation, and tissue repair in the intestine. Cell Host Microbe 12, 445–457 (2012).
Lee, M.W. et al. Activated type 2 innate lymphoid cells regulate beige fat biogenesis. Cell 160, 74–87 (2015).
Brestoff, J.R. et al. Group 2 innate lymphoid cells promote beiging of white adipose tissue and limit obesity. Nature 519, 242–246 (2015).
Oliphant, C.J. et al. MHCII-mediated dialog between group 2 innate lymphoid cells and CD4+ T cells potentiates type 2 immunity and promotes parasitic helminth expulsion. Immunity 41, 283–295 (2014).
Halim, T.Y. et al. Group 2 innate lymphoid cells are critical for the initiation of adaptive T helper 2 cell-mediated allergic lung inflammation. Immunity 40, 425–435 (2014).
Salimi, M. et al. A role for IL-25 and IL-33-driven type-2 innate lymphoid cells in atopic dermatitis. J. Exp. Med. 210, 2939–2950 (2013).
Kim, B.S. et al. TSLP elicits IL-33-independent innate lymphoid cell responses to promote skin inflammation. Sci. Transl. Med. 5, 170ra16 (2013).
Hams, E. et al. IL-25 and type 2 innate lymphoid cells induce pulmonary fibrosis. Proc. Natl. Acad. Sci. USA 111, 367–372 (2014).
McHedlidze, T. et al. Interleukin-33-dependent innate lymphoid cells mediate hepatic fibrosis. Immunity 39, 357–371 (2013).
Spits, H. et al. Innate lymphoid cells–a proposal for uniform nomenclature. Nat. Rev. Immunol. 13, 145–149 (2013).
Walker, J.A., Barlow, J.L. & McKenzie, A.N.J. Innate lymphoid cells–how did we miss them? Nat. Rev. Immunol. 13, 75–87 (2013).
Klose, C.S.N. et al. A T-bet gradient controls the fate and function of CCR6−RORγt+ innate lymphoid cells. Nature 494, 261–265 (2013).
Bernink, J.H. et al. Human type 1 innate lymphoid cells accumulate in inflamed mucosal tissues. Nat. Immunol. 14, 221–229 (2013).
Bernink, J.H. et al. Interleukin-12 and -23 control plasticity of CD127+ group 1 and group 3 innate lymphoid cells in the intestinal lamina propria. Immunity 43, 146–160 (2015).
Mjösberg, J.M. et al. Human IL-25- and IL-33-responsive type 2 innate lymphoid cells are defined by expression of CRTH2 and CD161. Nat. Immunol. 12, 1055–1062 (2011).
Garlanda, C., Dinarello, C.A. & Mantovani, A. The interleukin-1 family: back to the future. Immunity 39, 1003–1018 (2013).
Mortha, A. et al. Microbiota-dependent crosstalk between macrophages and ILC3 promotes intestinal homeostasis. Science 343, 1249288 (2014).
Monticelli, L.A. et al. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nat. Immunol. 12, 1045–1054 (2011).
Barlow, J.L. et al. IL-33 is more potent than IL-25 in provoking IL-13-producing nuocytes (type 2 innate lymphoid cells) and airway contraction. J. Allergy Clin. Immunol. 132, 933–941 (2013).
Hazenberg, M.D. & Spits, H. Human innate lymphoid cells. Blood 124, 700–709 (2014).
Munneke, J.M. et al. Activated innate lymphoid cells are associated with a reduced susceptibility to graft-versus-host disease. Blood 124, 812–821 (2014).
Suzukawa, M. et al. An IL-1 cytokine member, IL-33, induces human basophil activation via its ST2 receptor. J. Immunol. 181, 5981–5989 (2008).
Pecaric-Petkovic, T., Didichenko, S.A., Kaempfer, S., Spiegl, N. & Dahinden, C.A. Human basophils and eosinophils are the direct target leukocytes of the novel IL-1 family member IL-33. Blood 113, 1526–1534 (2009).
Halim, T.Y. et al. Retinoic-acid-receptor-related orphan nuclear receptor α is required for natural helper cell development and allergic inflammation. Immunity 37, 463–474 (2012).
Walker, J.A. et al. Bcl11b is essential for group 2 innate lymphoid cell development. J. Exp. Med. 212, 875–882 (2015).
Yu, Y. et al. The transcription factor Bcl11b is specifically expressed in group 2 innate lymphoid cells and is essential for their development. J. Exp. Med. 212, 865–874 (2015).
Califano, D. et al. Transcription factor Bcl11b controls identity and function of mature type 2 innate lymphoid cells. Immunity 43, 354–368 (2015).
Fuchs, A. et al. Intraepithelial type 1 innate lymphoid cells are a unique subset of IL-12- and IL-15-responsive IFN-γ-producing cells. Immunity 38, 769–781 (2013).
Wilson, C.B., Rowell, E. & Sekimata, M. Epigenetic control of T-helper-cell differentiation. Nat. Rev. Immunol. 9, 91–105 (2009).
Hoyler, T. et al. The transcription factor GATA-3 controls cell fate and maintenance of type 2 innate lymphoid cells. Immunity 37, 634–648 (2012).
Björklund, A.K. et al. The heterogeneity of human CD127+ innate lymphoid cells revealed by single-cell RNA sequencing. Nat. Immunol. 17, 451–460 (2016).
Yagi, R. et al. The transcription factor GATA3 is critical for the development of all IL-7Rα-expressing innate lymphoid cells. Immunity 40, 378–388 (2014).
Nussbaum, J.C. et al. Type 2 innate lymphoid cells control eosinophil homeostasis. Nature 502, 245–248 (2013).
Silver, J.S. et al. Inflammatory triggers associated with exacerbations of chronic obstructive pulmonary disease orchestrate plasticity of group 2 innate lymphoid cells in the lungs. Nat. Immunol. 10.1038/ni.3443 (2016).
Bal, S.M. et al. IL-1β, IL-4 and IL-12 control the fate of group 2 innate lymphoid cells in human airway inflammation. Nat. Immunol. 10.1038/ni.3444 (2016).
Sims, J.E. & Smith, D.E. The IL-1 family: regulators of immunity. Nat. Rev. Immunol. 10, 89–102 (2010).
Kearley, J. et al. Cigarette smoke silences innate lymphoid cell function and facilitates an exacerbated type I interleukin-33-dependent response to infection. Immunity 42, 566–579 (2015).
Turner, J.E. et al. IL-9-mediated survival of type 2 innate lymphoid cells promotes damage control in helminth-induced lung inflammation. J. Exp. Med. 210, 2951–2965 (2013).
Hepworth, M.R. et al. Innate lymphoid cells regulate CD4+ T-cell responses to intestinal commensal bacteria. Nature 498, 113–117 (2013).
Wenzel, S.E. Asthma phenotypes: the evolution from clinical to molecular approaches. Nat. Med. 18, 716–725 (2012).
Lambrecht, B.N. & Hammad, H. The immunology of asthma. Nat. Immunol. 16, 45–56 (2015).
Randolph, D.A., Stephens, R., Carruthers, C.J. & Chaplin, D.D. Cooperation between Th1 and Th2 cells in a murine model of eosinophilic airway inflammation. J. Clin. Invest. 104, 1021–1029 (1999).
Hansen, G., Berry, G., DeKruyff, R.H. & Umetsu, D.T. Allergen-specific Th1 cells fail to counterbalance Th2 cell-induced airway hyperreactivity but cause severe airway inflammation. J. Clin. Invest. 103, 175–183 (1999).
Nakao, F. et al. Association of IFN-gamma and IFN regulatory factor 1 polymorphisms with childhood atopic asthma. J. Allergy Clin. Immunol. 107, 499–504 (2001).
Yu, M. et al. Identification of an IFN-γ/mast cell axis in a mouse model of chronic asthma. J. Clin. Invest. 121, 3133–3143 (2011).
Sugimoto, T. et al. Interleukin 18 acts on memory T helper cells type 1 to induce airway inflammation and hyperresponsiveness in a naive host mouse. J. Exp. Med. 199, 535–545 (2004).
Soumelis, V. et al. Human epithelial cells trigger dendritic cell mediated allergic inflammation by producing TSLP. Nat. Immunol. 3, 673–680 (2002).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Anders, S., Pyl, P.T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome Res. 22, 1760–1774 (2012).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Roederer, M., Nozzi, J.L. & Nason, M.C. SPICE: exploration and analysis of post-cytometric complex multivariate datasets. Cytometry A 79, 167–174 (2011).
Kozuka, T., Sugita, M., Shetzline, S., Gewirtz, A.M. & Nakata, Y. c-Myb and GATA-3 cooperatively regulate IL-13 expression via conserved GATA-3 response element and recruit mixed lineage leukemia (MLL) for histone modification of the IL-13 locus. J. Immunol. 187, 5974–5982 (2011).
Acknowledgements
We thank H. Ueno, S. Hanabuchi, T. Kim and M. Ramaswamy for discussions; C. Harrod and C. Kiefer for critical reading of the manuscript; N. Loof, C. Boudreaux, K. Kayembe, C. Groves and R. Rayanki for support with cell sorting; H. Lu and A. Berlin for laboratory support; the LAR staff for maintaining the experimental animals; N. Baldwin for help in depositing RNA-sequencing data; and A. O'Bar and S. Zurawski for performing Luminex experiment. Supported by the Cancer Prevention and Research Institute of Texas (RP110319 (“Targeting Dendritic Cells to Block Immunosuppression in Breast Cancer”) to Y.-J.L.) and by the Japan Society for the Promotion of Science (KAKENHI Grant-in-Aid 23-10890 to Y.O.)
Author information
Authors and Affiliations
Contributions
Y.O. and Y.-J.L. conceived of the idea for this project; Y.O., J.S.S., L.T.-S., A.A.H. and Y.-J.L. designed experiments; Y.O., J.S.S., L.T.-S., M.A.C., J.P.B. and A.M.C. performed experiments and analyzed data; Y.O. and B.L.C. performed analysis of RNA sequencing data; and Y.O., J.S.S., L.T.-S., A.H.H. and Y.-J.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Y.O., J.S.S., A.M.C., A.A.H. and Y.-J.L. are employed by and shareholders of Medimmune.
Integrated supplementary information
Supplementary Figure 1 Identification of human ILC2s in peripheral blood and tonsils.
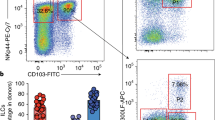
(a) List of cytokines tested for the ability to promote proliferation of ILC2s. (b) Expression of T-bet and GATA-3 in ILCs in PBMC examined by intra-nuclear transcription staining followed by flow cytometry. Subsets of ILCs were defined as in Fig. 1a. (c) Purity and yield of ILC2s from blood after cell sorting. (d) Purity and yield ILC2s after FACS sorting. (e) Schematic of identification of ILCs in human tonsils by flow cytometry. Antibodies against the following lineage makers (Lin) are used: CD3, CD4, CD14, CD16, CD19, CD20, CD34, CD123, TCRαβ, HLA-DR. ILC2s were identified as Lin- CD127+ CD56- CRTH2+. CD56brigtht NK cells were identified as Lin-CD127int c-Kit- CD56hi.
Supplementary Figure 2 IL-1β enhances the response of ILC2s to epithelial cytokines.
(a) Expression of CRLF2 in CD56brigtht NK cells, ILC2s and c-Kit+ CD127+ cells freshly isolated from peripheral blood or stimulated with IL-2 and IL-1β for 7 days. Expression of CRLF2 was normalized to GUSB and shown as arbitrary unit. (b) Cell numbers of ILC2s recovered after culturing with IL-2, IL-25 and IL-1β for 7 days (300 cells plated per well at day 0). (c) Scheme of the experiments comparing freshly isolated ILC2s and IL-1β-primed ILC2s. (d) Frequencies of phospho-NF-κB positive cells in Fig. 3f. (e) NF-κB phosphorylation in CD56brigtht NK cells, ILC2s and c-Kit+ CD127+ cells in freshly isolated from peripheral blood stimulated with IL-1β, IL-18 and IL-33 for 5 min. *P < 0.05, **P < 0.01 and *** P < 0.001 using one-way ANOVA followed by two-tailed t-test (a), one-way ANOVA followed by Dunnet’s test (b), or followed by Turkey’s test (d). Data are from 6 (a) and 5 (b) and 4 (d) independent experiments with independent donors or from one experiment representative of four (e) experiments with similar results. Data are represented as mean (± s.e.m.) (a,b).
Supplementary Figure 3 Validation of the expression changes in ILC2s stimulated with IL-1β.
(a) The surface expression of HLA-DR/DP/DQ, CD80 and CD40L measured by flow cytometry in ILC2s freshly isolated from peripheral blood or cultured with IL-2 and IL-1β for 7 days. (b) mRNA levels of GATA3, RORC and RORA in freshly isolated ILC2s, CD56bright NK cells and ILC2s stimulated with IL-2 and IL-1β for 5 days. Data are normalized to GUSB and shown as relative values and represented as mean (± s.e.m.). *P<0.05, ** P < 0.01, *** P < 0.001 analyzed using one-way ANOVA followed by Turkey’s test (b). Data are representative of 6 experiments with similar results (a), or 6 (b) experiments with independent donors.
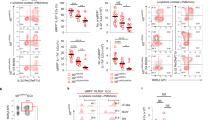
Supplementary Figure 4 IL-12 affects ILC2 functional gene expression and phenotype plasticity.
(a) Viabilities of ILC2s cultured with IL-2 and IL-1β with or without IL-12 for 7 days. (b) Schematic of identification of human c-Kit+ and c-Kit- subpopulations of ILC2s in human PBMC by flow cytometry. CD56brigtht NK cells and ILC2s were defined as in Fig. 1a and ILC2s were further divided by expression of c-Kit. Numbers adjacent to outlined areas indicate percent of parent population. (c) Recovery of c-Kit+ and c-Kit- ILC2s subpopulations from 150 ml whole blood. (d) Intracellular staining of IL-13, IL-5 and IFN-γ in c-Kit+ and c-Kit- ILC2s, and CD56brigtht NK cells cultured with or without IL-12 in the presence of IL-2 and IL-1β for 7 days followed by re-stimulation with PMA and ionomycin for 6 h in the presence of protein secretion inhibitors. (e) Frequencies of IL-5 and IL-13 double positive cells (left) and IFN-γ and IL-13 double positive cells (right) measured in d. (f) Expression of IL1RL1, IL17RB, CRLF2 and GATA3 in ILC2s freshly isolated from peripheral blood or cultured with IL-2 and IL-1β with or without IL-12 for 7 days. Data are normalized to GUSB and shown as relative values. * P < 0.05, **P < 0.01 and ***P < 0.001 as analyzed by two-tailed t-test (a), one-way ANOVA followed by Turkey’s test (e,f). Data are from 9 (a) or 6 (c,e,f) experiments with independent donors or from one experiment representative of 6 experiments (d) with similar results. Data are represented as mean (± s.e.m.) (f).
Supplementary Figure 5 IFNG and type-2-cytokine-encoding loci.
(a) IFNG locus and (b) Type 2 cytokine loci are shown with gray boxes depicting the distal regulatory regions for each locus. The red boxes approximately refer to the regions amplified by qPCR using specific primers as shown in Fig. 6. CNS, conserved non-coding sequences. CGRE, conserved GATA-3 response element. IE, IL4 intronic enhancer. HS, hyper sensitive site.
Supplementary Figure 6 Identification of ILC2 cells and NK cells in vivo and the effect of anti-IL-12 on IL-1-mediated ILC2 activation.
(a) The expression of lineage markers CD3, CD19, B220, CD5, TCRβ, TCRγδ, CD11c, F4/80, Gr1, Ter119, and CD27 on CD45+ cells, ILC2s and CD49b+ NKs measured by flow cytometry. (b) The concentration of IL-13 in the culture supernatant of ILC2 isolated from the lungs of the mice administered and stimulated as in Fig. 7g. Data are representative of 7 independent experiments (a). Data are represented as mean (b).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–4 (PDF 1150 kb)
Rights and permissions
About this article
Cite this article
Ohne, Y., Silver, J., Thompson-Snipes, L. et al. IL-1 is a critical regulator of group 2 innate lymphoid cell function and plasticity. Nat Immunol 17, 646–655 (2016). https://doi.org/10.1038/ni.3447
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3447
This article is cited by
-
A diversity of novel type-2 innate lymphoid cell subpopulations revealed during tumour expansion
Communications Biology (2024)
-
The modulation of pulmonary group 2 innate lymphoid cell function in asthma: from inflammatory mediators to environmental and metabolic factors
Experimental & Molecular Medicine (2023)
-
Interferons as negative regulators of ILC2s in allergic lung inflammation and respiratory viral infections
Journal of Molecular Medicine (2023)
-
Insights into the tumor microenvironment of B cell lymphoma
Journal of Experimental & Clinical Cancer Research (2022)
-
Innate lymphoid cells and cancer
Nature Immunology (2022)