Abstract
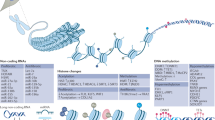
Rheumatoid arthritis synovial fibroblasts (RASFs) are the effector cells of cartilage and bone destruction. These cells show an 'intrinsically' activated and aggressive phenotype that results in the increased production of matrix-degrading enzymes and adhesion molecules, and is conserved over long-term passage in vitro. The three main mechanisms of epigenetic control—DNA methylation, histone modifications and microRNA activity—interact in the development of the RASF phenotype. The extent of global DNA methylation is reduced in synoviocytes in situ and RASFs in vitro. In addition, histone hyperacetylation occurs and specific microRNAs are expressed in RASFs. Normal synovial fibroblasts cultured in a hypomethylating milieu acquire an activated phenotype similar to that of RASFs. These findings suggest that epigenetic control, in particular the control of DNA methylation, is deficient in RASFs. Genome-wide analyses of the epigenome will enable the detection of additional genes involved in the pathogenesis of rheumatoid arthritis, the identification of epigenetic biomarkers, and potentially the development of a therapeutic regimen that targets activated RASFs.
Key Points
-
Rheumatoid arthritis synovial fibroblasts (RASFs) exhibit an aggressive phenotype that is characterized by the expression of matrix-degrading enzymes and adhesion molecules
-
The aggressive RASF phenotype can be induced in normal synovial fibroblasts by culture in a hypomethylating milieu
-
Histone modifications and microRNA binding have also been implicated in the development of RASF phenotype, which suggests that all three epigenetic pathways are altered in rheumatoid arthritis
-
Whole-epigenome analysis could lead to the discovery of novel genes and epigenetic biomarkers involved in the pathogenesis of rheumatoid arthritis, with potential implications for treatment
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Eggermann, T., Meyer, E., Caglayan, A. O., Dundar, M. & Schonherr, N. ICR1 epimutations in llp15 are restricted to patients with Silver–Russell syndrome features. J. Pediatr. Endocrinol. Metab. 21, 59–62 (2008).
Hauke, J. et al. Survival motor neuron gene 2 silencing by DNA methylation correlates with spinal muscular atrophy disease severity and can be bypassed by histone deacetylase inhibition. Hum. Mol. Genet. 18, 304–317 (2009).
Gerli, R. et al. Precocious atherosclerosis in rheumatoid arthritis: role of traditional and disease-related cardiovascular risk factors. Ann. NY Acad. Sci. 1108, 372–381 (2007).
Stöger, R. Epigenetics and obesity. Pharmacogenomics 9, 1851–1860 (2008).
Chiang, P. K., Lam, M. A. & Luo, Y. The many faces of amyloid β in Alzheimer's disease. Curr. Mol. Med. 8, 580–584 (2008).
Sokka, T. et al. Women, men, and rheumatoid arthritis: analyses of disease activity, disease characteristics, and treatments in the QUEST-RA Study. Arthritis Res. Ther. 11, R7 (2009).
Lu, Q. et al. Demethylation of CD40LG on the inactive X in T cells from women with lupus. J. Immunol. 179, 6352–6358 (2007).
Javierre, B. M., Esteller, M. & Ballestar, E. Epigenetic connections between autoimmune disorders and haematological malignancies. Trends Immunol. 29, 616–623 (2008).
Singal, R. & Ginder, G. D. DNA methylation. Blood 93, 4059–4070 (1999).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Turek-Plewa, J. & Jagodzinski, P. P. The role of mammalian DNA methyltransferases in the regulation of gene expression. Cell Mol. Biol. Lett. 10, 631–647 (2005).
Issa, J. P. CpG island methylator phenotype in cancer. Nat. Rev. Cancer 4, 988–993 (2004).
Ballestar, E. & Esteller, M. Methyl-CpG-binding proteins in cancer: blaming the DNA methylation messenger. Biochem. Cell Biol. 83, 374–384 (2005).
Robertson, K. D. DNA methylation and human disease. Nat. Rev. Genet. 6, 597–610 (2005).
Pountos, I., Corscadden, D., Emery, P. & Giannoudis, P. V. Mesenchymal stem cell tissue engineering: techniques for isolation, expansion and application. Injury 38 (Suppl. 4), S23–S33 (2007).
Boquest, A. C., Noer, A. & Collas, P. Epigenetic programming of mesenchymal stem cells from human adipose tissue. Stem Cell Rev. 2, 319–329 (2006).
Takada, I., Suzawa, M., Matsumoto, K. & Kato, S. Suppression of PPAR transactivation switches cell fate of bone marrow stem cells from adipocytes into osteoblasts. Ann. NY Acad. Sci. 1116, 182–195 (2007).
Nolte, S. V., Xu, W., Rennekampff, H. O. & Rodemann, H. P. Diversity of fibroblasts—a review on implications for skin tissue engineering. Cells Tissues Organs 187, 165–176 (2008).
Karouzakis, E., Neidhart, M., Gay, R. E. & Gay, S. Molecular and cellular basis of rheumatoid joint destruction. Immunol. Lett. 106, 8–13 (2006).
Bauer, S. et al. Fibroblast activation protein is expressed by rheumatoid myofibroblast-like synoviocytes. Arthritis Res. Ther. 8, R171 (2006).
Muller-Ladner, U., Pap, T., Gay, R. E., Neidhart, M. & Gay, S. Mechanisms of disease: the molecular and cellular basis of joint destruction in rheumatoid arthritis. Nat. Clin. Pract. Rheumatol. 1, 102–110 (2005).
Lafyatis, R. et al. Anchorage-independent growth of synoviocytes from arthritic and normal joints. Stimulation by exogenous platelet-derived growth factor and inhibition by transforming growth factor-β and retinoids. J. Clin. Invest. 83, 1267–1276 (1989).
Konttinen, Y. T. et al. Analysis of 16 different matrix metalloproteinases (MMP-1 to MMP-20) in the synovial membrane: different profiles in trauma and rheumatoid arthritis. Ann. Rheum. Dis. 58, 691–697 (1999).
Rinaldi, N. et al. Differential expression and functional behaviour of the αv and β3 integrin subunits in cytokine-stimulated fibroblast-like cells derived from synovial tissue of rheumatoid arthritis and osteoarthritis in vitro. Ann. Rheum. Dis. 56, 729–736 (1997).
Firestein, G. S., Alvaro-Gracia, J. M. & Maki, R. Quantitative analysis of cytokine gene expression in rheumatoid arthritis. J. Immunol. 144, 3347–3353 (1990).
Huber, L. C. et al. Histone deacetylase/acetylase activity in total synovial tissue derived from rheumatoid arthritis and osteoarthritis patients. Arthritis Rheum. 56, 1087–1093 (2007).
Stanczyk, J. et al. Altered expression of µRNA in synovial fibroblasts and synovial tissue in rheumatoid arthritis. Arthritis Rheum. 58, 1001–1009 (2008).
Neidhart, M. et al. Retrotransposable L1 elements expressed in rheumatoid arthritis synovial tissue: association with genomic DNA hypomethylation and influence on gene expression. Arthritis Rheum. 43, 2634–2647 (2000).
Ali, M. et al. Overexpression of transcripts containing LINE-1 in the synovia of patients with rheumatoid arthritis. Ann. Rheum. Dis. 62, 663–666 (2003).
Kuchen, S. et al. The L1 retroelement-related p40 protein induces p38δ MAP kinase. Autoimmunity 37, 57–65 (2004).
Karouzakis, E. et al. Genomic hypomethylation of rheumatoid arthritis synovial fibroblasts [abstract 745]. Arthritis Rheum. 56 (Suppl.), S317 (2007).
Schulz, W. A., Steinhoff, C. & Florl, A. R. Methylation of endogenous human retroelements in health and disease. Curr. Top. Microbiol. Immunol. 310, 211–250 (2006).
Nile, C. J., Read, R. C., Akil, M., Duff, G. W. & Wilson, A. G. Methylation status of a single CpG site in the IL6 promoter is related to IL-6 messenger RNA levels and rheumatoid arthritis. Arthritis Rheum. 58, 2686–2693 (2008).
Kimura, F. et al. Decrease of DNA methyltransferase 1 expression relative to cell proliferation in transitional cell carcinoma. Int. J. Cancer 104, 568–578 (2003).
Karouzakis, E., Gay, R. E., Kolling, C., Gay, S. & Neidhart, M. Epigenetic profile of gene expression in normal and rheumatoid arthritis synovial fibroblasts [abstract 939]. Arthritis Rheum. 58 (Suppl.), S514 (2008).
Xu, G. L. et al. Chromosome instability and immunodeficiency syndrome caused by mutations in a DNA methyltransferase gene. Nature 402, 187–191 (1999).
Karouzakis, E., Gay, R. E., Michel, B. A., Gay, S. & Neidhart, M. The increased expression of β1 integrin (CD29) in rheumatoid synovial fibroblasts is due to a milieu favouring genomic hypomethylation [abstract OP-0110]. Ann. Rheum. Dis. 67 (Suppl. II), 84 (2008).
Hmadcha, A., Bedoya, F. J., Sobrino, F. & Pintado, E. Methylation-dependent gene silencing induced by interleukin 1β via nitric oxide production. J. Exp. Med. 190, 1595–1604 (1999).
Wehbe, H., Henson, R., Meng, F., Mize-Berge, J. & Patel, T. Interleukin-6 contributes to growth in cholangiocarcinoma cells by aberrant promoter methylation and gene expression. Cancer Res. 66, 10517–10524 (2006).
Hodge, D. R. et al. IL-6 enhances the nuclear translocation of DNA cytosine-5-methyltransferase 1 (DNMT1) via phosphorylation of the nuclear localization sequence by Akt kinase. Cancer Genomics Proteomics 4, 387–398 (2007).
Chuang, L. S. et al. Human DNA-(cytosine-5)-methyltransferase–PCNA complex as a target for p21waf1. Science 277, 1996–2000 (1997).
Szyf, M. The role of DNA methyltransferase 1 in growth control. Front. Biosci. 6, D599–D609 (2001).
Deng, J. & Szyf, M. Downregulation of DNA (cytosine-5)-methyltransferase is a late event in NGF-induced PC12 cell differentiation. Brain Res. Mol. Brain Res. 71, 23–31 (1999).
Esteller, M. Epigenetics in cancer. N. Engl. J. Med. 358, 1148–1159 (2008).
Takami, N. et al. Hypermethylated promoter region of DR3, the death receptor 3 gene, in rheumatoid arthritis synovial cells. Arthritis Rheum. 54, 779–787 (2006).
Karouzakis, E., Gay, R. E., Kolling, C., Gay, S. & Neidhart, M. Epigenetic approaches to the study of the pathogenesis of rheumatic disease. Eur. Musculoskeletal Rev. 3, 41–43 (2008).
Clark, S. J., Statham, A., Stirzaker, C., Molloy, P. L. & Frommer, M. DNA methylation: bisulphite modification and analysis. Nat. Protoc. 1, 2353–2364 (2006).
Tost, J. & Gut, I. G. DNA methylation analysis by pyrosequencing. Nat. Protoc. 2, 2265–2275 (2007).
Holemon, H. et al. MethylScreen: DNA methylation density monitoring using quantitative PCR. Biotechniques 43, 683–693 (2007).
Weber, M. et al. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat. Genet. 39, 457–466 (2007).
Mulero-Navarro, S. & Esteller, M. Epigenetic biomarkers for human cancer: the time is now. Crit. Rev. Oncol. Hematol. 68, 1–11 (2008).
Butcher, L. M. & Beck, S. Future impact of integrated high-throughput methylome analyses on human health and disease. J. Genet. Genomics 35, 391–401 (2008).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Karouzakis, E., Gay, R., Gay, S. et al. Epigenetic control in rheumatoid arthritis synovial fibroblasts. Nat Rev Rheumatol 5, 266–272 (2009). https://doi.org/10.1038/nrrheum.2009.55
Issue Date:
DOI: https://doi.org/10.1038/nrrheum.2009.55
This article is cited by
-
Dimethyl Fumarate Inhibits Fibroblast Like Synoviocytes-mediated Inflammation and Joint Destruction in Rheumatoid Arthritis
Inflammation (2023)
-
Epigenome association study for DNA methylation biomarkers in buccal and monocyte cells for female rheumatoid arthritis
Scientific Reports (2021)
-
Penta-acetyl Geniposide Suppresses Migration, Invasion, and Inflammation of TNF-α-Stimulated Rheumatoid Arthritis Fibroblast-Like Synoviocytes Involving Wnt/β-Catenin Signaling Pathway
Inflammation (2021)
-
NR1D1 modulates synovial inflammation and bone destruction in rheumatoid arthritis
Cell Death & Disease (2020)
-
miR-449a inhibits cell proliferation, migration, and inflammation by regulating high-mobility group box protein 1 and forms a mutual inhibition loop with Yin Yang 1 in rheumatoid arthritis fibroblast-like synoviocytes
Arthritis Research & Therapy (2019)